Benjamin Good
Job Title
Assistant Professor
Employer
The Scripps Research Institute

Review details about the recently announced changes to study and work permits that apply to master’s and doctoral degree students. Read more
The Doctor of Philosophy in Bioinformatics (PhD)is an interdisciplinary program that combines the application of computer technology to the management and analysis of biological data. The result is that computers are being used to organize data generated from experiments into databases, develop new algorithms and software, and use this software for the interpretation and analysis of the data into meaningful biological information. For the past ten years, our PhD program has been training students to organize, visualize, analyze and interpret biological data. Students have access to world renowned bioinformaticians at the University of British Columbia, Simon Fraser University and the BC Cancer Agency, and have exposure to the latest technologies to develop their skills.
Strategic Program Objectives:
The Bioinformatics PhD program integrates academic centres in computer science, statistics, molecular biology and biotechnology, with translational groups at hospitals and at the clinical interface. The innovative partnership among the University of British Columbia, Simon Fraser University and the BC Cancer Agency allows students' access to experts in the field of bioinformatics, and exposure to original research and opportunities to complete significant practical work on real bioinformatics problems. Internships allow student mobility between Canadian and international universities, institutions and industries to further enhance collaborations among Canadian high-technology research groups in both the private and public sectors.
The major requirement for the Ph.D. is completion of a research dissertation meeting the Faculty of Graduate and Postdoctoral Studies requirements. There are no specific course requirements for the Ph.D. degree program apart from the dissertation. However, the student's Ph.D. dissertation committee has the prerogative to impose course requirements where course deficiencies are perceived.
All doctoral students are required to successfully complete a comprehensive examination, which consists of an oral and written component within the first 36 months of study. All students are required to present a Bioinformatics graduate program seminar upon completion of their program, and before their dissertation defense.
A student's committee for the doctorate will consist of the dissertation supervisor and three others. The supervisor and at least one other member must be members of the Bioinformatics graduate program.
Students must secure a supervisor before they can be admitted into the program. As well, they must meet the minimum admission requirements set out by Graduate and Post-doctoral Studies at UBC.
The Faculty of Graduate and Postdoctoral Studies establishes the minimum admission requirements common to all applicants, usually a minimum overall average in the B+ range (76% at UBC). The graduate program that you are applying to may have additional requirements. Please review the specific requirements for applicants with credentials from institutions in:
Each program may set higher academic minimum requirements. Please review the program website carefully to understand the program requirements. Meeting the minimum requirements does not guarantee admission as it is a competitive process.
Applicants from a university outside Canada in which English is not the primary language of instruction must provide results of an English language proficiency examination as part of their application. Tests must have been taken within the last 24 months at the time of submission of your application.
Minimum requirements for the two most common English language proficiency tests to apply to this program are listed below:
Overall score requirement: 100
Reading
22
Writing
21
Speaking
21
Listening
22
Overall score requirement: 7.0
Reading
6.0
Writing
6.0
Speaking
6.0
Listening
6.0
Some programs require additional test scores such as the Graduate Record Examination (GRE) or the Graduate Management Test (GMAT). The requirements for this program are:
The GRE is not required.
Students admitted to the Ph.D. degree program normally possess an M.Sc. degree in Bioinformatics or a related area, with clear evidence of research ability or potential.
CV, Official transcripts, three letters of reference, Official English exam scores (if required)
All applicants have to submit transcripts from all past post-secondary study. Document submission requirements depend on whether your institution of study is within Canada or outside of Canada.
Many programs require a statement of interest, sometimes called a "statement of intent", "description of research interests" or something similar.
Students in research-based programs usually require a faculty member to function as their thesis supervisor. Please follow the instructions provided by each program whether applicants should contact faculty members.
Permanent Residents of Canada must provide a clear photocopy of both sides of the Permanent Resident card.
All applicants must complete an online application form and pay the application fee to be considered for admission to UBC.
Students who secure an NSERC-CREATE scholarship will undertake a 3-4 month internship that may be local, within Canada or at an international University or Institution.
Bioinformatics faculty are spread throughout the UBC campus, as well as off-campus at the BC Cancer Research Centre or hospital research labs and Institutions.
| Fees | Canadian Citizen / Permanent Resident / Refugee / Diplomat | International |
|---|---|---|
| Application Fee | $118.50 | $168.25 |
| Tuition * | ||
| Installments per year | 3 | 3 |
| Tuition per installment | $1,875.34 | $3,294.66 |
| Tuition per year (plus annual increase, usually 2%-5%) | $5,626.02 | $9,883.98 |
| Int. Tuition Award (ITA) per year (if eligible) | $3,200.00 (-) | |
| Other Fees and Costs | ||
| Student Fees (yearly) | $1,144.10 (approx.) | |
| Costs of living | Estimate your costs of living with our interactive tool in order to start developing a financial plan for your graduate studies. | |
Applicants to UBC have access to a variety of funding options, including merit-based (i.e. based on your academic performance) and need-based (i.e. based on your financial situation) opportunities.
All students accepted by a faculty member and enrolled in the program will be paid a minimum stipend of $24,300/year.
This results in a net balance (any funding provided to the student minus tuition and fees) mean of $28,191 and median of $29,590.
All applicants are encouraged to review the awards listing to identify potential opportunities to fund their graduate education. The database lists merit-based scholarships and awards and allows for filtering by various criteria, such as domestic vs. international or degree level.
Many professors are able to provide Research Assistantships (GRA) from their research grants to support full-time graduate students studying under their supervision. The duties constitute part of the student's graduate degree requirements. A Graduate Research Assistantship is considered a form of fellowship for a period of graduate study and is therefore not covered by a collective agreement. Stipends vary widely, and are dependent on the field of study and the type of research grant from which the assistantship is being funded.
Graduate programs may have Teaching Assistantships available for registered full-time graduate students. Full teaching assistantships involve 12 hours work per week in preparation, lecturing, or laboratory instruction although many graduate programs offer partial TA appointments at less than 12 hours per week. Teaching assistantship rates are set by collective bargaining between the University and the Teaching Assistants' Union.
Academic Assistantships are employment opportunities to perform work that is relevant to the university or to an individual faculty member, but not to support the student’s graduate research and thesis. Wages are considered regular earnings and when paid monthly, include vacation pay.
Canadian and US applicants may qualify for governmental loans to finance their studies. Please review eligibility and types of loans.
All students may be able to access private sector or bank loans.
UBC has working agreements with MPower Financing - an organization providing international students with no-cosigner, no-collateral education loans to study in Canada - and Windmill Microlending - an organization providing loans to skilled immigrants.
Many foreign governments provide support to their citizens in pursuing education abroad. International applicants should check the various governmental resources in their home country, such as the Department of Education, for available scholarships.
The possibility to pursue work to supplement income may depend on the demands the program has on students. It should be carefully weighed if work leads to prolonged program durations or whether work placements can be meaningfully embedded into a program.
International students enrolled as full-time students with a valid study permit can work on campus for unlimited hours and work off-campus for no more than 24 hours a week during academic sessions.
A good starting point to explore student jobs is the UBC Work Learn program or a Co-Op placement.
Students with taxable income in Canada may be able to claim federal or provincial tax credits.
Canadian residents with RRSP accounts may be able to use the Lifelong Learning Plan (LLP) which allows students to withdraw amounts from their registered retirement savings plan (RRSPs) to finance full-time training or education for themselves or their partner.
Please review Filing taxes in Canada on the student services website for more information.
Applicants have access to the cost estimator to develop a financial plan that takes into account various income sources and expenses.
12 students graduated between 2005 and 2013. Of these, career information was obtained for 12 alumni (based on research conducted between Feb-May 2016):
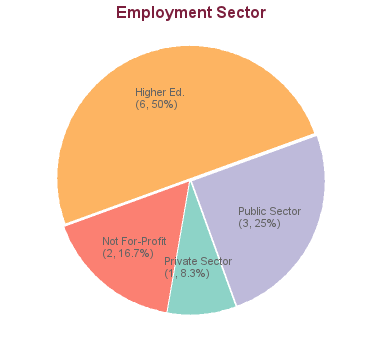
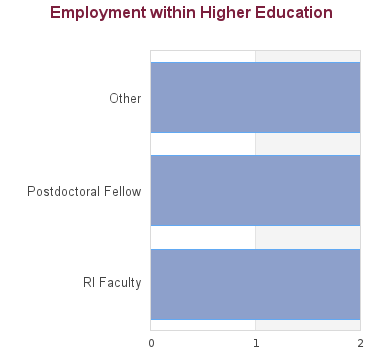
As biological datasets increase exponentially in both size and complexity, bioinformatics tools have central importance in fields and industries ranging from environmental management, forestry, aquaculture, and biofuels to personalized medicine, drug development, preventative medicine and gene therapy. Individuals who can analyuze and interpret large data sets or "big data" are highly sought after by both public and private sector employers. Academic positions at Universities are widely available in all fields of study.
Ph.D. graduates from the program have gone on to pursue post-doctoral studies at Stanford, Harvard school of Medicine, Max Delbruck Centre in Berlin, Broad Institute at MIT and Harvard, Ontario Cancer Institute, Howard Hughes Medical Institute in Santa Cruz, and locally at UBC and SFU. One graduate is an assistant professor at Simon Fraser University and another is an assistant professor at the University of Dalhousie in Halifax, NS.
These statistics show data for the Doctor of Philosophy in Bioinformatics (PhD). Data are separated for each degree program combination. You may view data for other degree options in the respective program profile.
| 2024 | 2023 | 2022 | 2021 | 2020 | |
|---|---|---|---|---|---|
| Applications | 33 | 24 | 22 | 36 | 33 |
| Offers | 8 | 5 | 8 | 10 | 9 |
| New Registrations | 4 | 2 | 5 | 8 | 8 |
| Total Enrolment | 48 | 52 | 49 | 47 | 44 |
Students in research-based programs usually require a faculty member to function as their thesis supervisor. Please follow the instructions provided by each program whether applicants should contact faculty members.
These videos contain some general advice from faculty across UBC on finding and reaching out to a supervisor. They are not program specific.
This list shows faculty members with full supervisory privileges who are affiliated with this program. It is not a comprehensive list of all potential supervisors as faculty from other programs or faculty members without full supervisory privileges can request approvals to supervise graduate students in this program.
Bioinformatics combines computational and biological disciplines.
Departments/Programs may update graduate degree program details through the Faculty & Staff portal. To update contact details for application inquiries, please use this form.
